Most of the studies that have used S9.6 to identify R-loops in eukaryotes have not used RNase A to eliminate any artifacts due to free RNA, which might be present during cell lysis and/or harvest of the nucleic acid. Recent work has shown that a single-chain variable domain of the S9.6 antibody can bind RNA:RNA duplexes with an affinity that is only 5.6-fold weaker than to RNA:DNA duplexes, raising the serious concern that S9.6 can indeed cross react with RNA species. This multi-antibody complex would reflect the combination of affinities of multiple antibodies. Such ELISA measurements can be complicated by multiple antibodies binding to a single long duplex. While S9.6 antibody can recognize RNA:DNA duplexes, complete characterization of the binding specificity of S9.6 was initially limited to ELISA measurements on its binding to long nucleic acid duplexes. Therefore, these well-documented regions with R-loops are ideal positive control targets of IP using S9.6 antibody. Our original description of kilobase long mammalian genomic R-loops was further built upon and had the advantage of several lines of independent evidence including (a) the large body of IgH switch DNA sequence and recombination junctional sequence information (b) many functional studies of IgH switch region transcription (c) concurrent studies of IgH switch region orientation and (d) detailed in vitro biochemical studies of transcription through switch regions. Moreover, we did not observe any large effects of parallel RNase A treatment in vivo or in vitro. This genomic R-loop analysis was based on (a) bisulfite sequencing and (b) oligonucleotide hybridization (colony lift hybridization), and both were done with and without an RNase H challenge treatment to confirm the RNA:DNA conformation. Physical presence of long genomic R-loops in eukaryotes was first described at the IgH switch regions. Caution should be used when interpreting S9.6 data, and confirmation by independent structural and functional methods is essential. ConclusionĪny use of the S9.6 antibody must be preceded by RNase A treatment to remove free ssRNA that may compete for the S9.6 binding by forming RNA:RNA regions or short, transient RNA:DNA duplexes. With RNase A treatment, a signal can be detected over background, but only within a limited 2 or 3-fold range, even with a stable kilobase-long genomic R-loop. Without RNase A treatment, known regions of R-loop formation containing RNA:DNA duplexes can not be reliably detected. 
We find that optimal detection of RNA:DNA duplexes requires removal of ssRNA using RNase A. The RNase treatments included RNase H to destroy the RNA in an RNA:DNA duplex and RNase A to destroy single-stranded (ss) RNA to prevent it from binding S9.6 directly (as duplex RNA) and to prevent the ssRNA from annealing to the genome, resulting in adventitious RNA:DNA hybrids. We tested IP using S9.6 with and without various RNase treatments. The R-loops at this locus can be induced by using cytokines to stimulate transcription from germline transcript promoters. FindingsĪs our test locus, we chose the most well-documented site for kilobase-long mammalian genomic R-loops, the immunoglobulin heavy chain locus (IgH) class switch regions. Here we investigate whether RNase A is needed to obtain reliable IP with S9.6. Fold back of ssRNA can readily generate RNA:RNA duplexes that may bind the S9.6 antibody, and adventitious binding of RNA may also create short RNA:DNA regions. Most IP protocols do not pre-clear the genomic nucleic acid with RNase A to remove free RNA. However, recent work has demonstrated that a variable domain of S9.6 binds AU-rich RNA:RNA duplexes with a K D that is only 5.6-fold weaker than for RNA:DNA duplexes.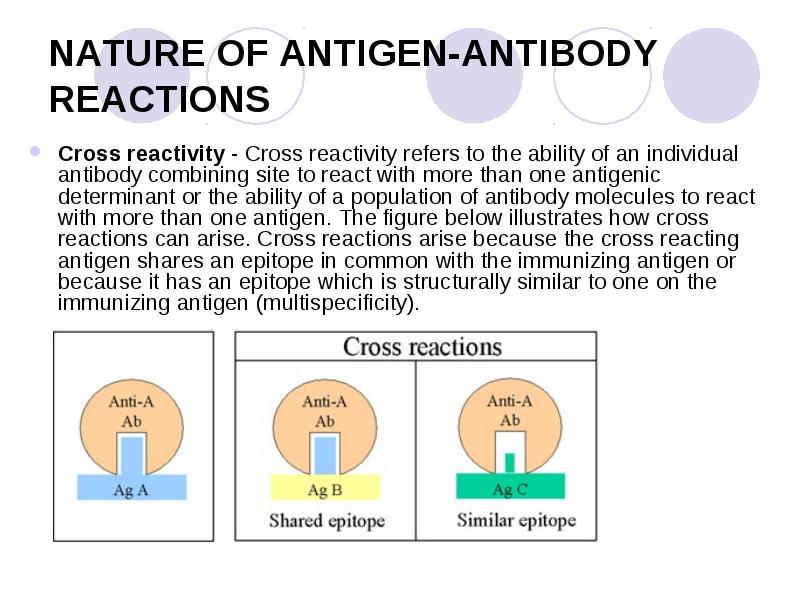  
A mouse monoclonal antibody called S9.6 has been used for immunoprecipitation (IP) to identify R-loops, based on the assumption that it is specific for RNA:DNA over other nucleic acid duplexes. Long genomic R-loops in eukaryotes were first described at the immunoglobulin heavy chain locus switch regions using bisulfite sequencing and functional studies. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
